 |
highres2standard_warp_inv is validconvert_xfm -omat highres2example_func.mat -inverse example_func2highres.mat
invwarp --ref==highres --warp=highres2standard_warp --out=highres2standard_warp_inv
and the output mask still has problem.
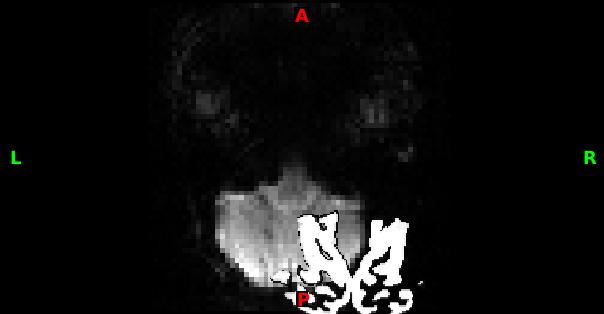
Hi Vasudev,No - your functional and structural data should be in the resolution they were acquired in respectively, and should match the resolution of the images you used in the registration process. The dimensions you have quoted look like reasonable values for T1/functional EPI data.To solve your problem, you need to identify the specific point of failure, and focus on fixing that. Refer to the list of steps in my previous email.In your other email thread, it looked to me like the non-linear registration was not ideal - this could be caused by a poor brain extraction, or an atypical brain. In your most recent screenshot it looks like something more fundamental has gone wrong - I would hazard a guess that one of the files you have used is incorrect.PaulOn Feb 20 2020, at 9:43 am, Dev vasu <[log in to unmask]> wrote:Dear Sir,- that all of the input files have the correct dimensions , is it necessary that dimensions of all functional and structural files should be same ?The Dimensions of all structural files issizeof_hdr 348data_type FLOAT32dim0 3dim1 160dim2 256dim3 256dim4 1dim5 1dim6 1dim7 1vox_units mmtime_units sdatatype 16nbyper 4bitpix 32pixdim0 0.000000pixdim1 1.000000pixdim2 1.000000pixdim3 1.000000pixdim4 1.000000pixdim5 0.000000pixdim6 0.000000pixdim7 0.000000vox_offset 352Dimensions of all Functional filessizeof_hdr 348data_type INT16dim0 4dim1 64dim2 64dim3 46dim4 300dim5 1dim6 1dim7 1vox_units mmtime_units sdatatype 4nbyper 2bitpix 16pixdim0 0.000000pixdim1 3.000000pixdim2 3.000000pixdim3 3.000000pixdim4 2.500000pixdim5 1.000000pixdim6 0.253332pixdim7 47455.246094vox_offset 352cal_max 0.0000cal_min 0.0000scl_slope 1.000000scl_inter 0.000000phase_dim 0freq_dim 0slice_dim 3slice_name alternating_increasing_2slice_code 5slice_start 0slice_end 45slice_duration 0.000000time_offset 0.000000intent Unknownintent_code 0intent_nameintent_p1 0.000000intent_p2 0.000000intent_p3 0.000000qform_name Scanner Anatqform_code 1On Thu, Feb 20, 2020 at 10:26 AM paul mccarthy <[log in to unmask]> wrote:Hi Vasudev,As I said in your other thread, this is due to a poor registration. Double check the following:- that all of the input files have the correct dimensions- The structural brain extraction is good- The func-struc registration is good- The struc-standard registration is good- The process you used to generatehighres2standard_warp_invis validPaulOn Feb 19 2020, at 8:09 pm, Dev vasu <[log in to unmask]> wrote:Dear all,I have done functional preprocessing using FEAT and i would like to perform seed based correlation for visual cortex seed regions, my seed is defined in MNI152 space.The seed and example_func.nii.gz are completely mismatched ( see picture ) , I have used Visual_thr_mask.nii.gz as seed ( see attachment) and tried to transform by doingapplywarp -i Visual_thr_mask.nii.gz -r reg/example_func.nii.gz -o Visual_func--postmat=reg/highres2example_func.mat -w reg/highres2standard_warp_invI dont understand what is the problem with my approach-could you please provideme some input.To unsubscribe from the FSL list, click the following link:To unsubscribe from the FSL list, click the following link:To unsubscribe from the FSL list, click the following link:
To unsubscribe from the FSL list, click the following link:
https://www.jiscmail.ac.uk/cgi-bin/webadmin?SUBED1=FSL&A=1
To unsubscribe from the FSL list, click the following link:
https://www.jiscmail.ac.uk/cgi-bin/webadmin?SUBED1=FSL&A=1